DeepLesion: a large-scale and diverse CT lesion dataset
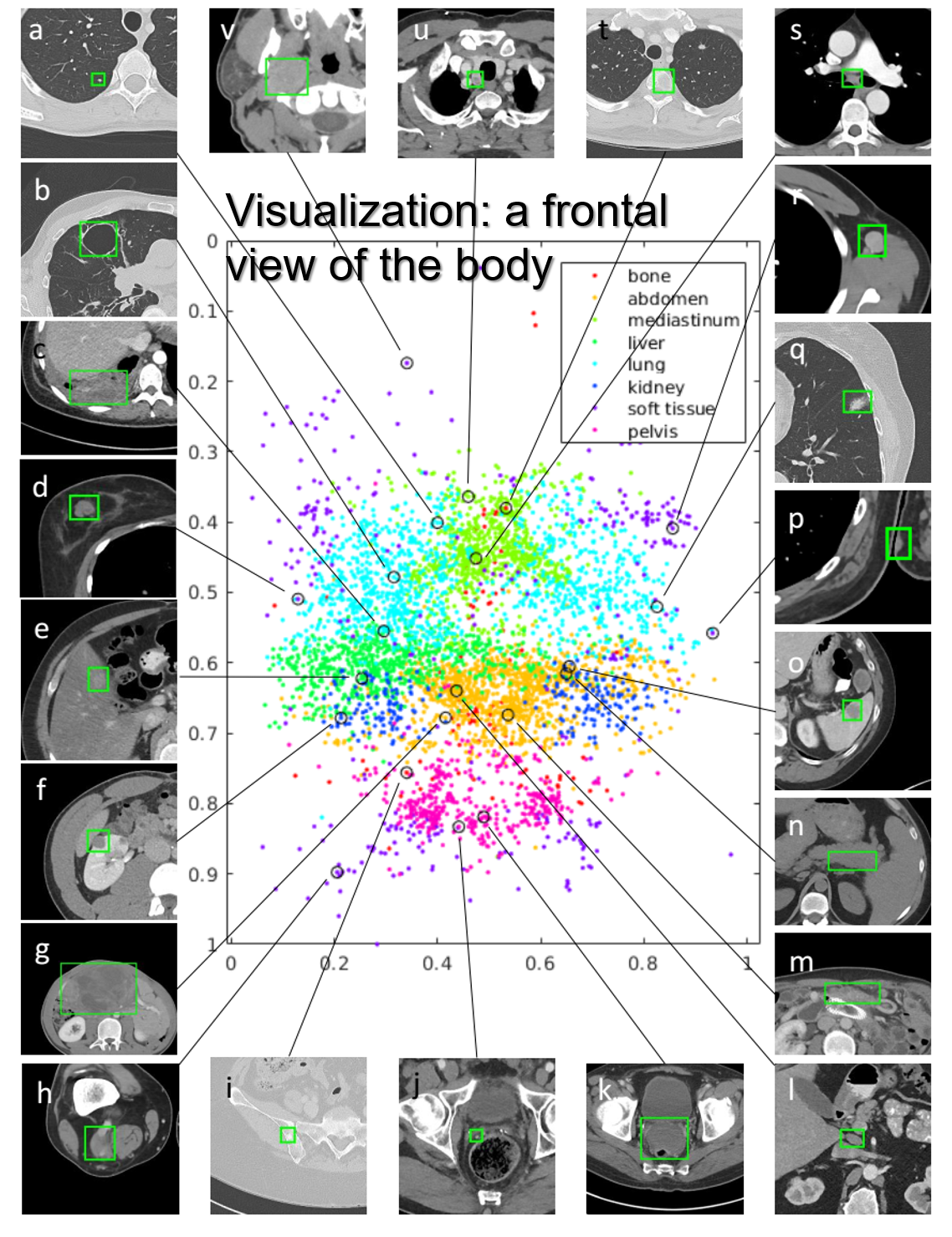
DeepLesion is a large-scale and diverse database of lesions in CT images. It was collected based on the bookmarks in the picture archiving and communication system of NIH. Currently, it contains 32,735 lesions from 32,120 axial CT slices of 4,427 patients. It can be used in studies of multi-class lesion detection, retrieval, segmentation, etc.
DeepLesion has been released. Download it without any paperwork here: https://nihcc.app.box.com/v/DeepLesion.
Please refer to our paper:
- DeepLesion: Automated Mining of Large-Scale Lesion Annotations and Universal Lesion Detection with Deep Learning (paper) Ke Yan, Xiaosong Wang, Le Lu, Ronald M. Summers, Journal of Medical Imaging, 2018.7.
- Deep Lesion Graphs in the Wild: Relationship Learning and Organization of Significant Radiology Image Findings in a Diverse Large-scale Lesion Database (paper, supplementary) Ke Yan, Xiaosong Wang, Le Lu, Ling Zhang, Adam P. Harrison, Mohammadhadi Bagheri, and Ronald M. Summers, IEEE CVPR, 2018.6.
Papers using DeepLesion:
- Lesion detection
- Ke Yan et al., "3D Context Enhanced Region-based Convolutional Neural Network for End-to-End Lesion Detection," MICCAI, 2018 (pdf, code)
- Youbao Tang et al., "ULDor: A Universal Lesion Detector for CT Scans with Pseudo Masks and Hard Negative Example Mining," ISBI, 2019 (pdf)
- Xudong Wang et al., "Towards Universal Object Detection by Domain Attention", CVPR, 2019 (pdf)
- Zihao Li et al., "MVP-Net: Multi-view FPN with Position-aware Attention for Deep Universal Lesion Detection", MICCAI, 2019
- Martin Zlocha et al., "Improving RetinaNet for CT Lesion Detection with Dense Masks from Weak RECIST Labels", MICCAI, 2019
- Qingyi Tao et al., "Improving Deep Lesion Detection Using 3D Contextual and Spatial Attention", MICCAI, 2019
- Lesion segmentation
- Jinzheng Cai et al., "Accurate Weakly-Supervised Deep Lesion Segmentation using Large-Scale Clinical Annotations: Slice-Propagated 3D Mask Generation from 2D RECIST," MICCAI, 2018 (pdf)
- Lesion measurement
- Youbao Tang et al., "Semi-Automatic RECIST Labeling on CT Scans with Cascaded Convolutional Neural Networks," MICCAI, 2018 (pdf)
- Lesion classification
- Ke Yan et al., "Holistic and Comprehensive Annotation of Clinically Significant Findings on Diverse CT Images: Learning from Radiology Reports and Label Ontology," CVPR, 2019 (pdf, code+model)
- Multi-task lesion analysis
- Ke Yan, et al., "MULAN: Multitask Universal Lesion Analysis Network for Joint Lesion Detection, Tagging, and Segmentation," MICCAI, 2019 (pdf, code+model)
- Lesion retrieval and matching
- Ke Yan et al., "Deep Lesion Graphs in the Wild: Relationship Learning and Organization of Significant Radiology Image Findings in a Diverse Large-scale Lesion Database," CVPR, 2018 (pdf)
- Image processing
- Youbao Tang et al., "CT Image Enhancement Using Stacked Generative Adversarial Networks and Transfer Learning for Lesion Segmentation Improvement," MLMI, 2018 (pdf)
- Wei-An Lin et al., "DuDoNet: Dual Domain Network for CT Metal Artifact Reduction," CVPR, 2019
If you are not familiar with medical image analysis, it is recommended to take a look at our lesion classification code for basic techniques such as loading annotations and 3D data, intensity windowing of 16-bit png files, pixel spacing normalization, etc.
