Self-supervised Learning of Pixel-wise Anatomical Embeddings in Radiological Images (arXiv)
Ke Yan, Jinzheng Cai, Dakai Jin, Shun Miao, Adam P. Harrison, Dazhou Guo, Youbao Tang, Jing Xiao, Jingjing Lu, Le Lu
arXiv, 2020.11.
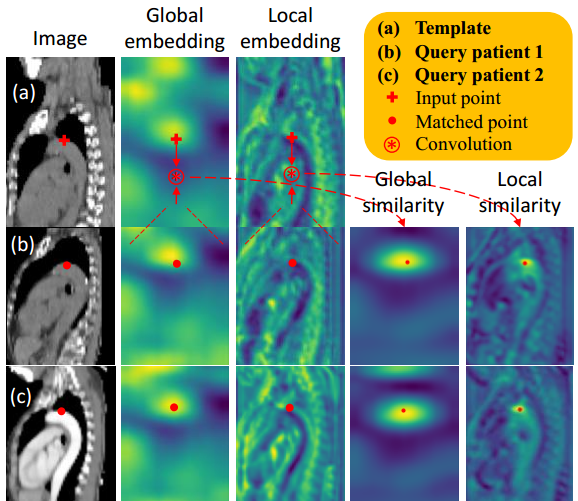
Self-supervised Anatomical eMbedding (SAM) is introduced in this paper. SAM generates semantic embeddings for each image pixel in radiological images (CT, X-ray, etc.) that describes its anatomical location. SAM is unsupervised and easy to train. It can be used for one-shot landmark detection and lesion matching, outperforming widely-used registration algorithms while being 200 times faster.
Learning from Multiple Datasets with Heterogeneous and Partial Labels for Universal Lesion Detection in CT (arXiv)
Ke Yan, Jinzheng Cai, Youjing Zheng, Adam P. Harrison, Dakai Jin, You-Bao Tang, Yu-Xing Tang, Lingyun Huang, Jing Xiao, Le Lu
IEEE Trans. on Medical Imaging, 2020.9.
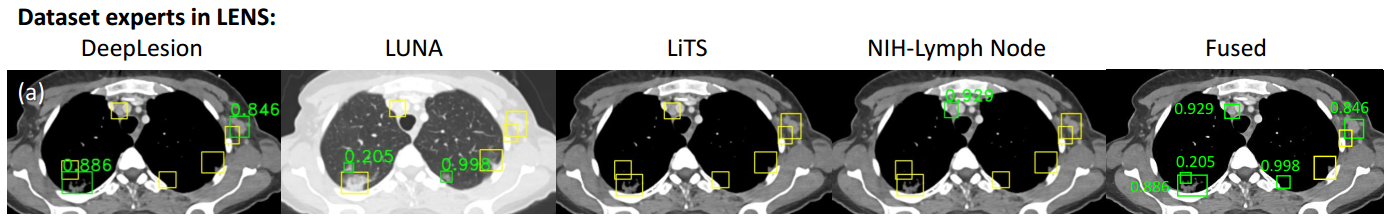
We design Lesion ENSemble (LENS) to learn from multiple heterogeneous lesion datasets for improved universal lesion detection. We also propose strategies to mine missing lesion annotations from partially-labeled datasets by exploiting clinical prior knowledge and cross-dataset knowledge transfer.
MULAN: Multitask Universal Lesion Analysis Network for Joint Lesion Detection, Tagging, and Segmentation (arXiv, code+model)
Ke Yan, Youbao Tang, Yifan Peng, Veit Sandfort, Mohammadhadi Bagheri, Zhiyong Lu, Ronald M. Summers
MICCAI, 2019.11.
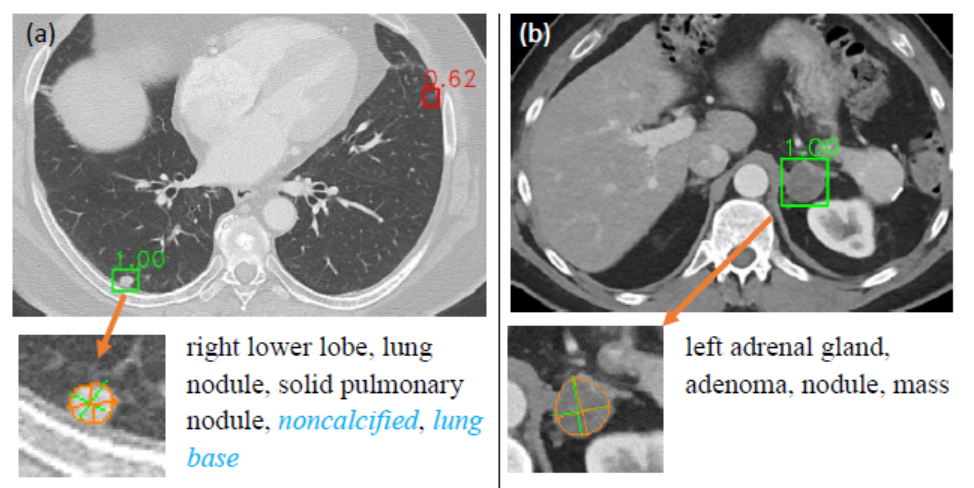
We propose multitask universal lesion analysis network (MULAN) for joint detection, tagging, and segmentation of a variety of lesions in CT images. It is a useful tool for both clinicians and CAD researchers.
Holistic and Comprehensive Annotation of Clinically Significant Findings on Diverse CT Images: Learning from Radiology Reports and Label Ontology (arXiv, code+label)
Ke Yan, Yifan Peng, Veit Sandfort, Mohammadhadi Bagheri, Zhiyong Lu, Ronald M. Summers
IEEE CVPR, 2019, oral presentation, 2019.6.
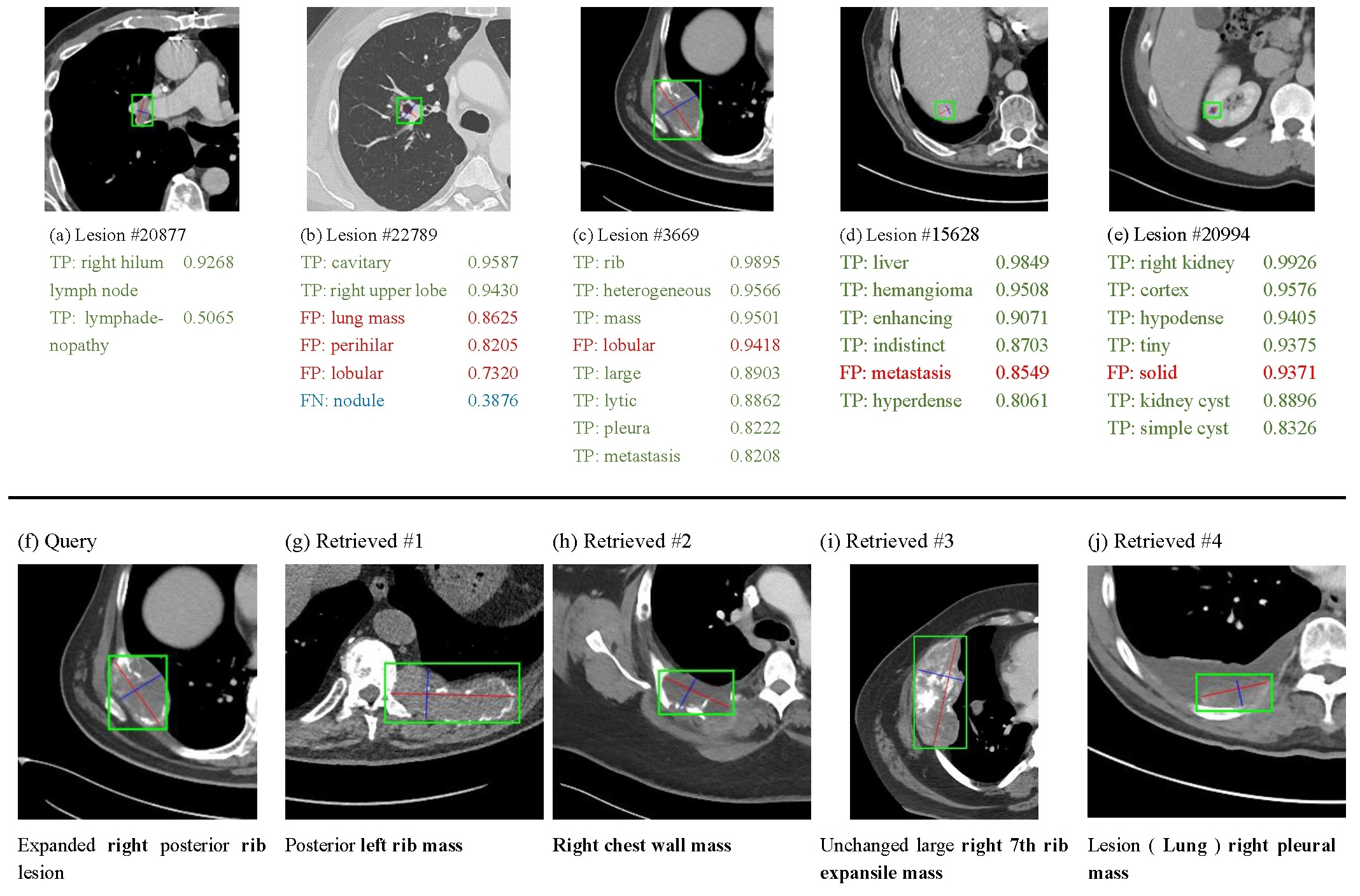
We propose Lesion Annotation Network (LesaNet) to predict multiple labels (body part, type, attributes) for a lesion image. The labels are learned from radiology reports and a lesion ontology. LesaNet can also be used for lesion retrieval.
Deep Lesion Graphs in the Wild: Relationship Learning and Organization of Significant Radiology Image Findings in a Diverse Large-scale Lesion Database (dataset, paper, supplementary)
Ke Yan, Xiaosong Wang, Le Lu, Ling Zhang, Adam P. Harrison, Mohammadhadi Bagheri, and Ronald M. Summers
IEEE CVPR, 2018.6.
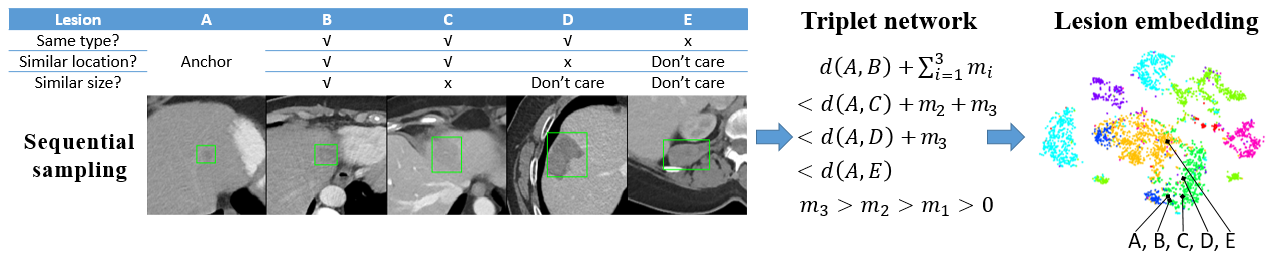
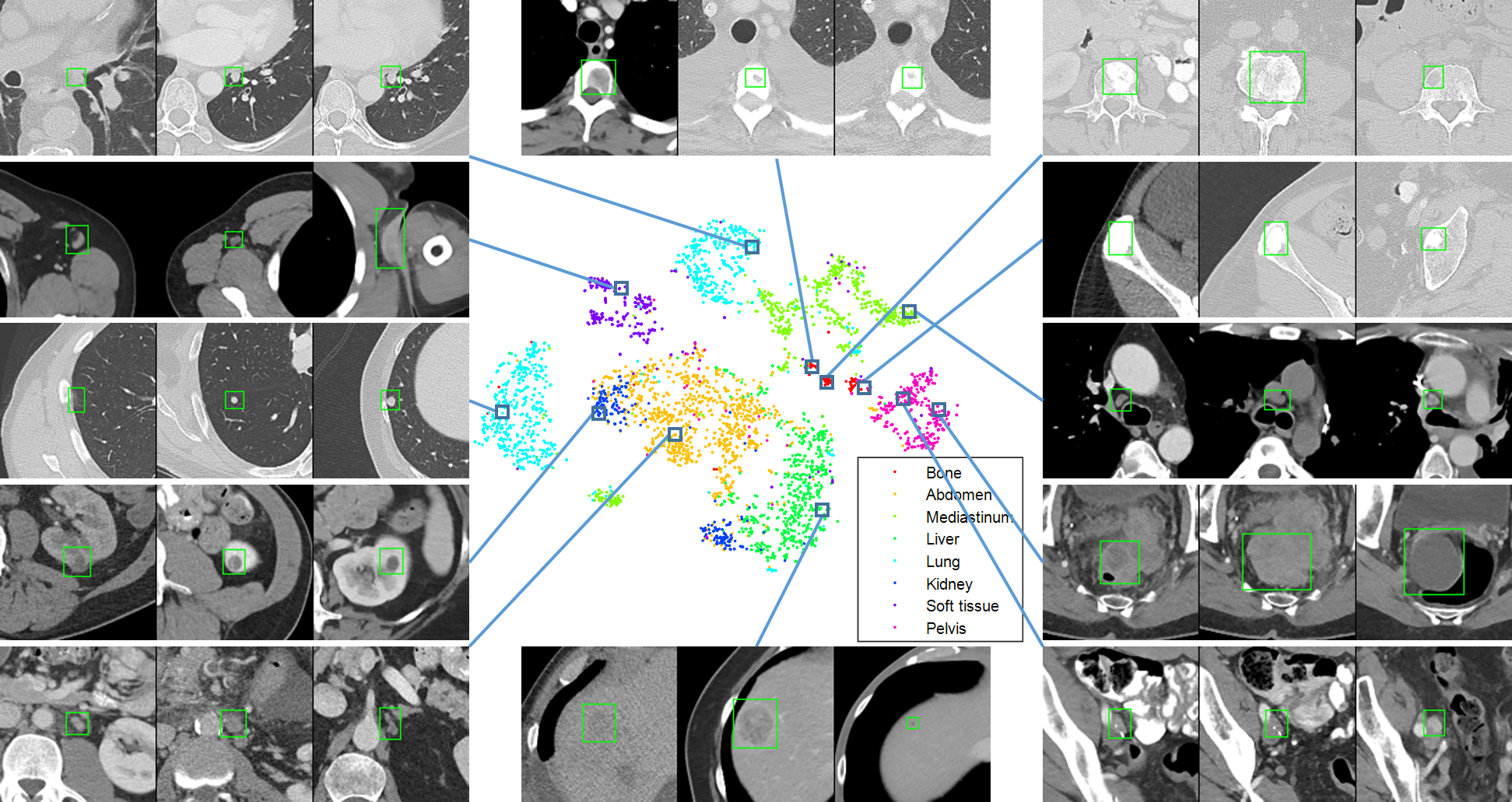
We present DeepLesion, a large-scale and diverse database of lesions in CT images. It can be used in studies of lesion detection, retrieval, segmentation, etc. A triplet network is proposed to learn lesion embeddings from weak cues for content-based lesion retrieval and intra-patient lesion matching.
3D Context Enhanced Region-based Convolutional Neural Network for End-to-End Lesion Detection (arXiv, code)
Ke Yan, Mohammadhadi Bagheri, and Ronald M. Summers
International Conference on Medical Image Computing & Computer Assisted Intervention (MICCAI), 2018.9.
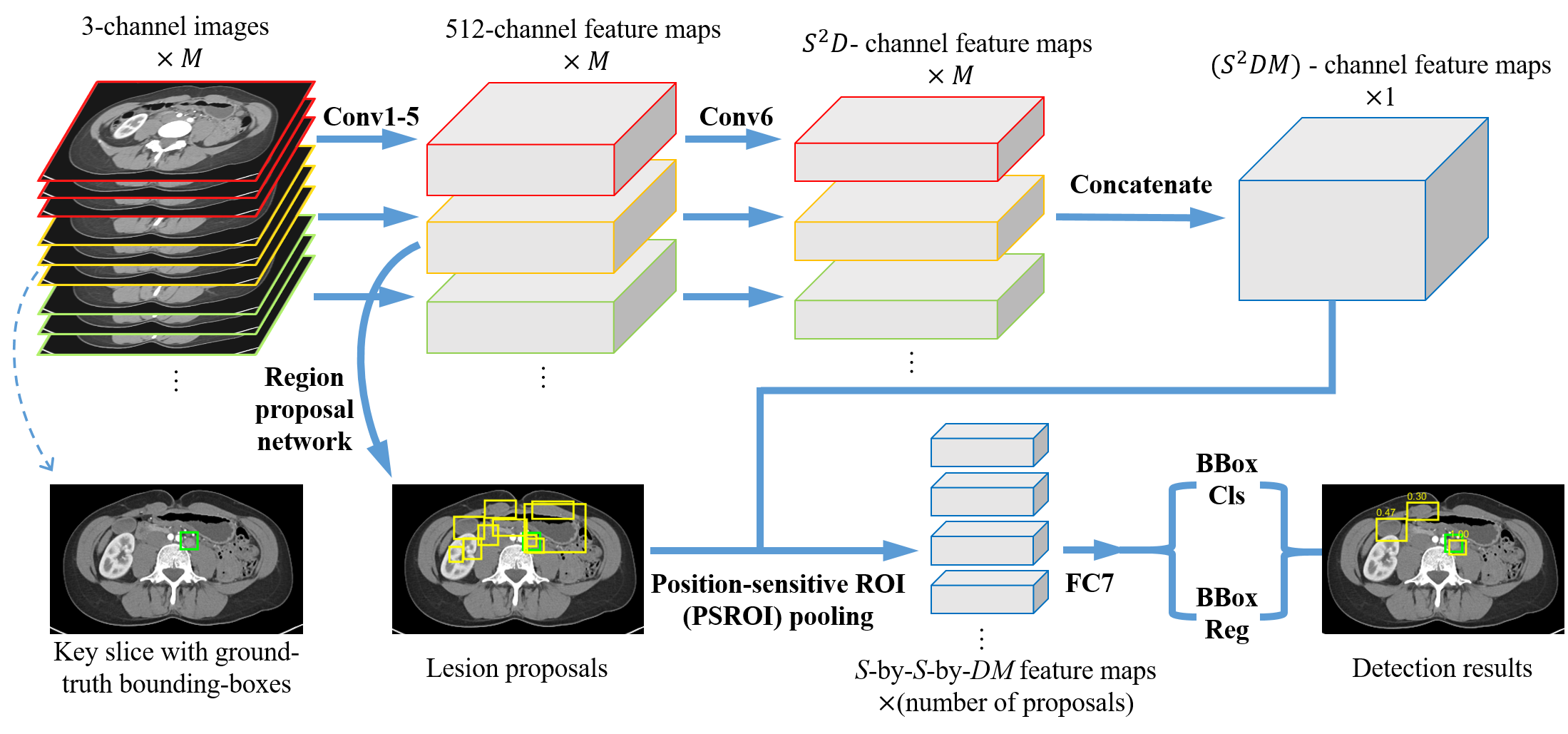
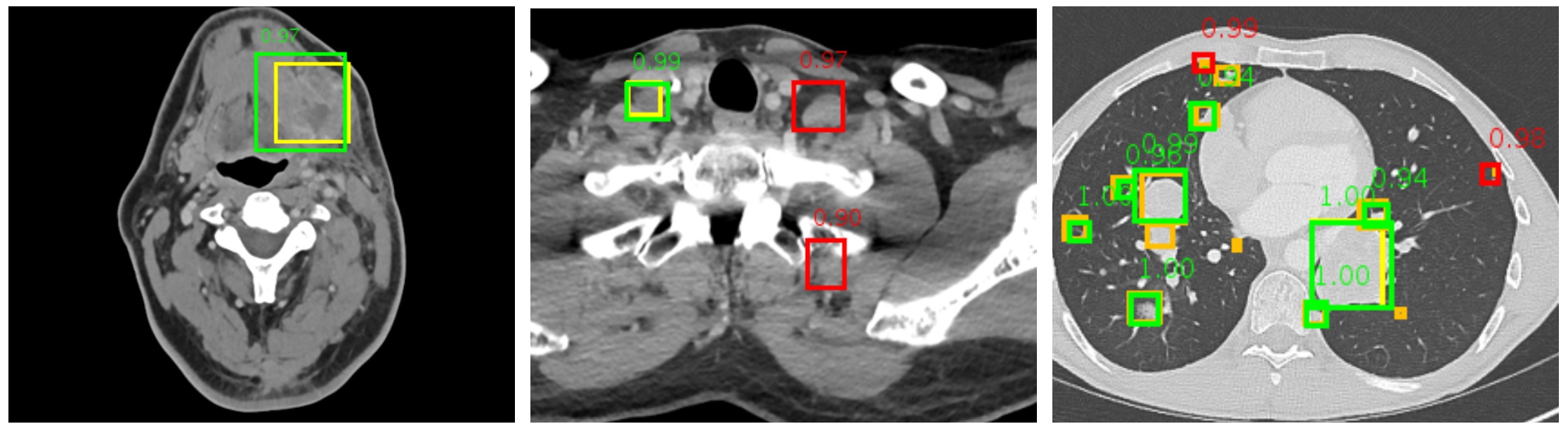
We present 3D context enhanced region-based CNN (3DCE) to leverage the 3D context when detecting lesions in volumetric data. 3DCE is memory-friendly, end-to-end, and simple to implement and train. Based on DeepLesion, we also develop a universal lesion detector that can help radiologists find all types of lesions with one unified framework.
Unsupervised Body Part Regression via Spatially Self-ordering Convolutional Neural Networks (arXiv, code))
Ke Yan, Le Lu, and Ronald M. Summers
IEEE ISBI (oral), 2018.4.
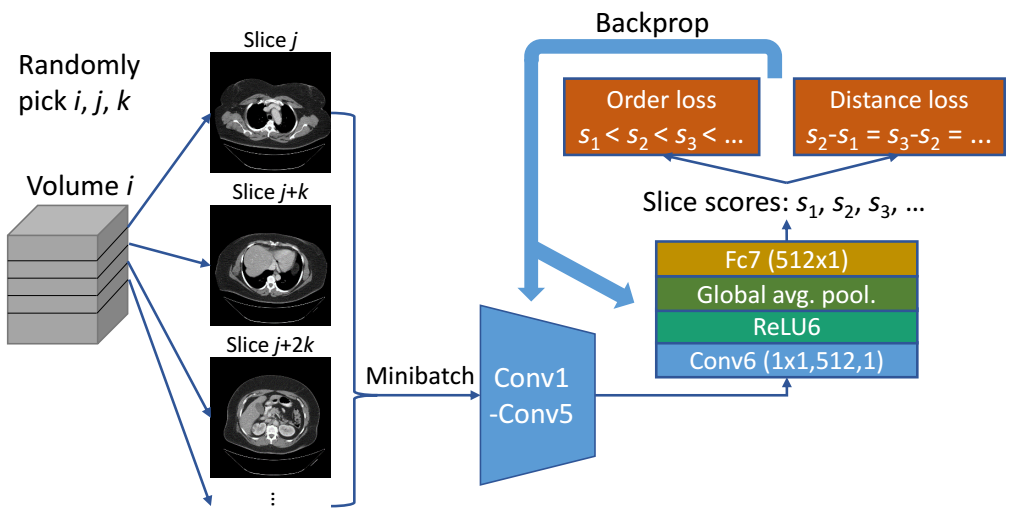
We propose unsupervised body part regressor (UBR), which builds a coordinate system for the body and outputs a continuous score for each slice. The score represents the normalized position of the body part in the slice.
DeepLesion: Automated Deep Mining, Categorization and Detection of Significant Radiology Image Findings using Large-Scale Clinical Lesion Annotations (arXiv)
Ke Yan*, Xiaosong Wang*, Le Lu, and Ronald M. Summers
RSNA (scientific poster), 2017.11.
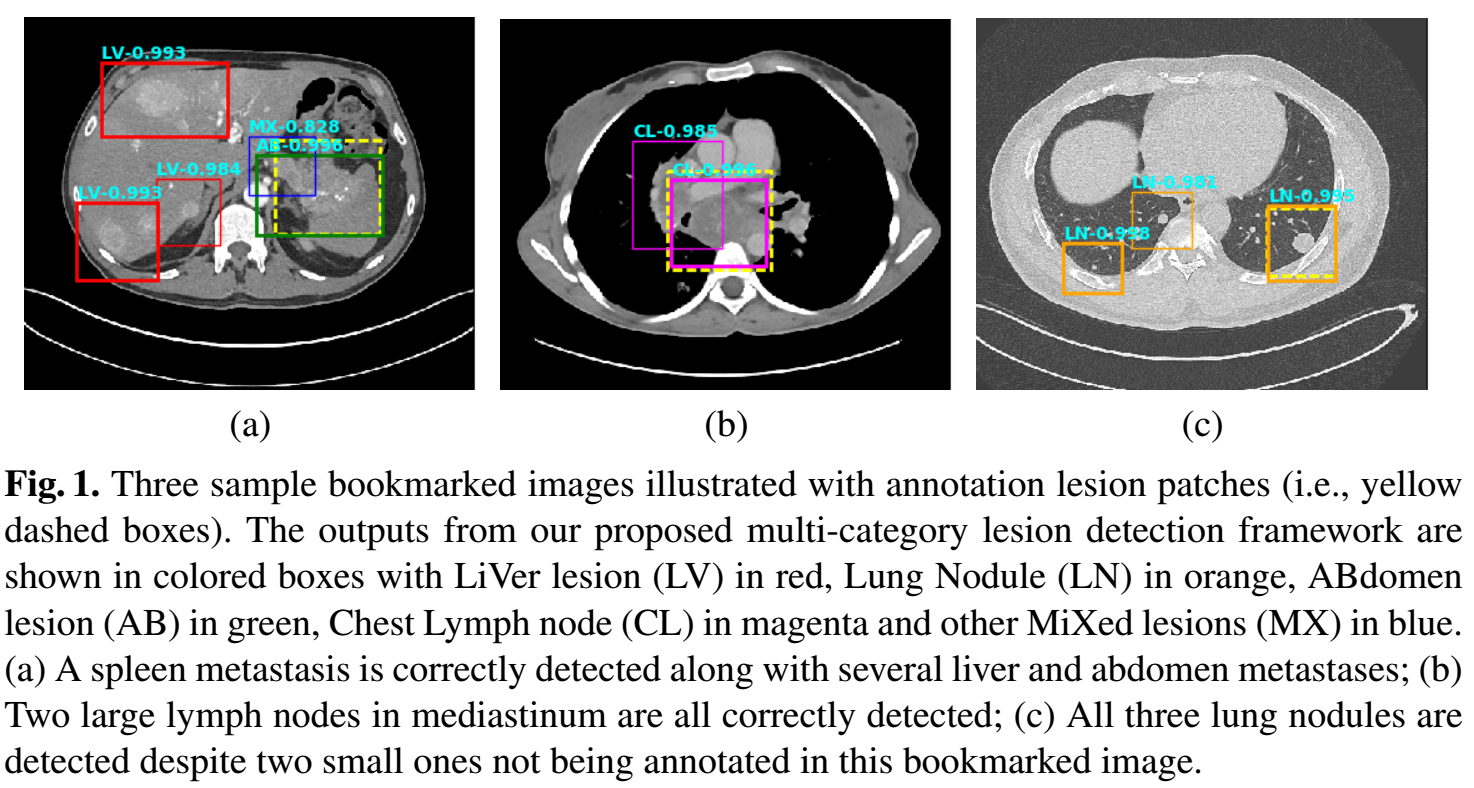
Lesion annotations are mined from PACS. A lesion detection algorithm based on Faster RCNN and an unsupervised categorization framework is proposed.
Learning classification and regression models for data with drift based on transfer samples (pdf, Matlab code)
Ke Yan, David Zhang, and Yong Xu
IEEE Trans. on Instrumentation and Measurement (IF = 2.456), 2017.
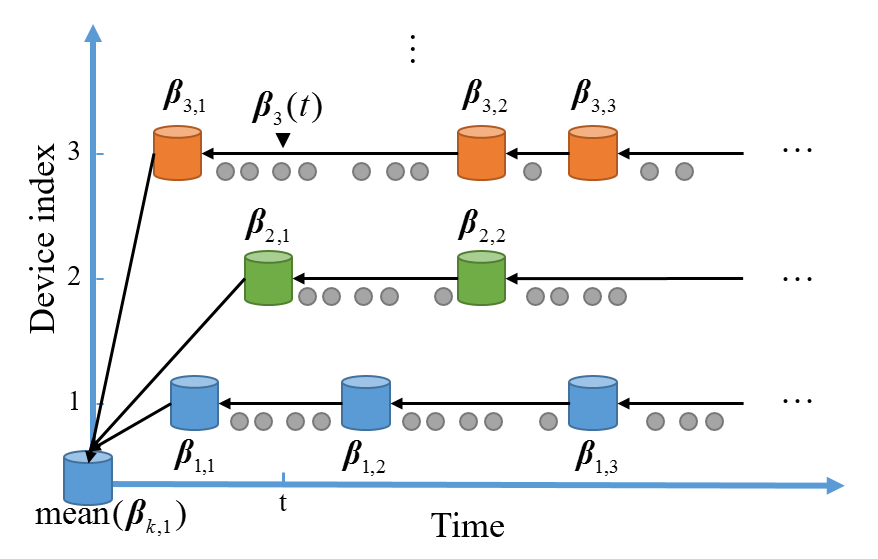
We aim to compensate the two largest influential factors in e-nose applications: instrumental variation and time-varying drift. The transfer-sample-based multitask learning (TMTL) algorithm defines each device or time period as a domain, then learns a prediction model for each domain. Two paradigms, parallel and serial transfer, are designed according to the prior knowledge of the two influential factors. A dynamic model strategy is also proposed to handle continuous sample stream in the MTL framework.
Learning Domain-Invariant Subspace Using Domain Features and Independence Maximization (pdf, Matlab code, domain adaptation toolbox)
Ke Yan, Lu Kou, and David Zhang
IEEE Trans. on Cybernetics (IF = 4.943), 2017.
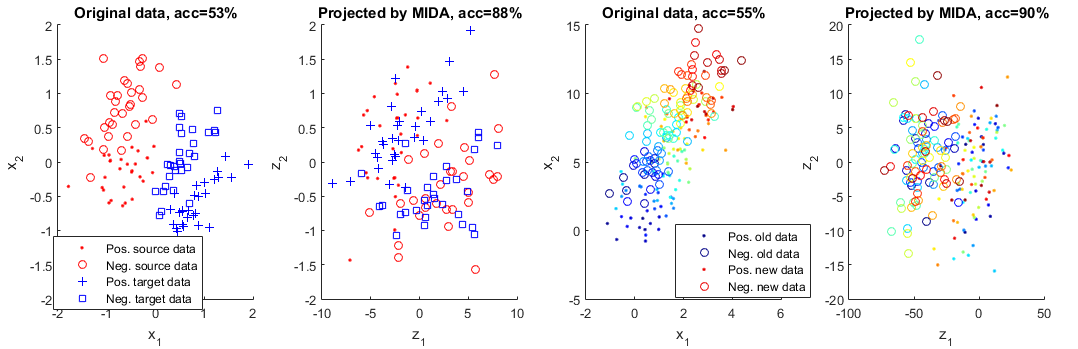
Maximum independence domain adaptation (MIDA) is proposed to learn a domain-invariant subspace. It can be applied in all kinds of domain adaptation problems, including discrete or continuous distributional change, supervised/semi-supervised/unsupervised, multiple domains, classification or regression, etc. We first design "domain features", then maximize the independence between them and the learned features.
Correcting instrumental variation and time-varying drift: a transfer learning approach with autoencoders (pdf, Python code)
Ke Yan and David Zhang
IEEE Trans. on Instrumentation and Measurement (IF = 1.790), 2016.
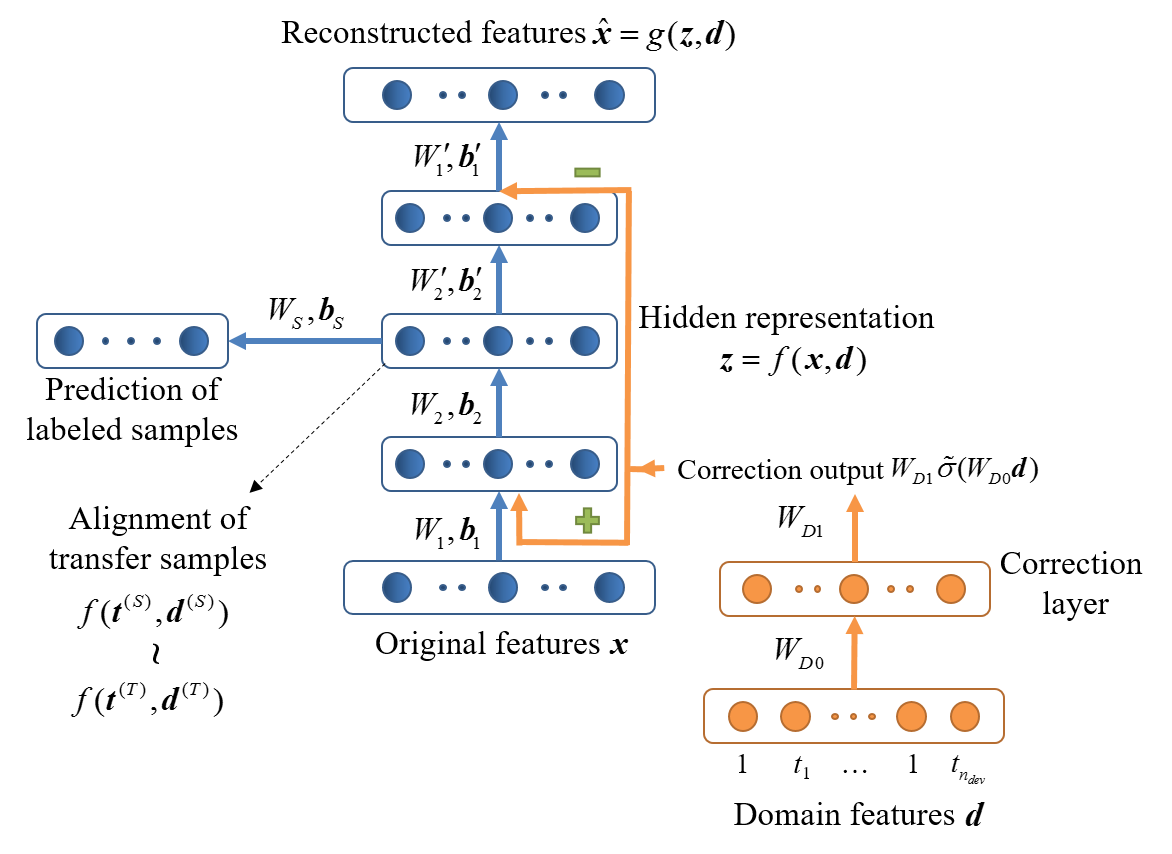
Drift correction autoencoder (DCAE) is proposed to correct instrumental variation and time-varying drift. We enhance the structure and objective function of the original autoencoder, so that the two factors can be explicitly modeled and compensated. A drift-corrected and discriminative representation of the original data is thus learned.
Calibration transfer and drift compensation of e-noses via coupled task learning (pdf, Matlab code)
Ke Yan and David Zhang
Sensors and Actuators B: Chemical (IF = 4.097), 2015.
The preliminary version of TMTL with only 2 tasks. The key idea is to align the transfer samples at the model level.
Improving the transfer ability of prediction models for electronic noses (pdf, Matlab code)
Ke Yan and David Zhang
Sensors and Actuators B: Chemical (IF = 4.097), 2015.
A novel variable standardization method and a regularization strategy, windowed piecewise direct standardization (WPDS) and standardization-error-based model improvement (SEMI), are proposed to deal with instrumental variation.
Feature selection and analysis on correlated gas sensor data with recursive feature elimination (pdf, Matlab code)
Ke Yan and David Zhang
Sensors and Actuators B: Chemical (IF = 4.097), 2015.
Introducing correlation bias reduction (CBR), an improvement to the feature selection algorithm SVM-RFE for correlated features.
Design of a breath analysis system for diabetes diagnosis and blood glucose level prediction (pdf)
Ke Yan, David Zhang, Darong Wu, Hua Wei, and Guangming Lu
IEEE Trans. on Biomedical Engineering (IF = 2.347), 2014.
The hardware and software of an e-nose-based system are introduced.
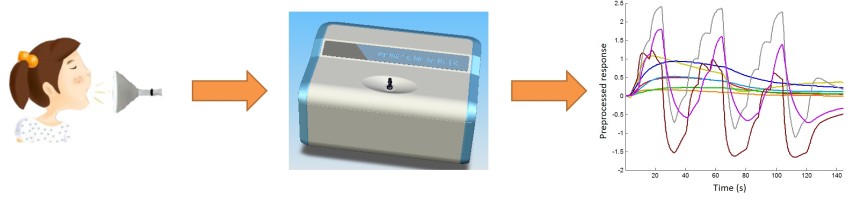
Blood glucose prediction by breath analysis system with feature selection and model fusion (pdf)
Ke Yan and David Zhang
36th Annual Intl. Conf. of the IEEE Engineering in Medicine and Biology Society (EMBC), Chicago, 2014, oral presentation
Predicting the blood glucose of 203 breath samples, mean relative absolute error = 20.69%.
Sensor evaluation in a breath analysis system (pdf)
Ke Yan and David Zhang
Intl. Conf. Medical Biometrics (ICMB), Shenzhen, 2014
A novel breath analysis system for diabetes diagnosis (pdf)
Ke Yan and David Zhang
Intl. Conf. Computerized Healthcare (ICMB), Hong Kong, 2012
Gabor Surface Feature for face recognition (pdf, Matlab code, OpenCV code)
Ke Yan, Youbin Chen and David Zhang
1st Asian Conference on Pattern Recognition (ACPR), Beijing, 2011
An improvement to the Gabor+LBP feature. Good performance on FERET dataset.
Patent: An exhaled gas detection system (in Chinese), Pending
一种呼出气体的检测系统 (link)
张大鹏, 闫轲, 卢光明
公开中, 公开号 CN103245705 A